In data analysis using R, the need to convert character columns to factors is common. Character columns often contain categorical data, and converting them to factors enables R to interpret and analyze the data more effectively. Factors represent categorical variables with distinct levels, aiding in statistical modeling (e.g., ANOVA, MANOVA) and visualization.
Data type conversion is a fundamental aspect of data preprocessing, influencing the success of subsequent analyses. While we previously learned how to convert multiple columns to numeric in R with dplyr and base functions, this post will cover two distinct approaches to converting all character columns to factors. Specifically, we will explore two methods to use R to convert all character columns to factors — 1) using base functions and 2) using the dplyr mutate() function.
Table of Contents
- Outline
- Prerequisites
- Data Types
- Synthetic Data
- Convert All Character Columns to Factor with Base R
- Convert All Character Columns to Factor in R with dplyr
- Comparing Methods for Changing Multiple Character Columns to Factors in R
- Conclusion
- Resources
Outline
This post is structured as follows: First, we get into the prerequisites, ensuring you have a foundational understanding of data types in R. Next, we generate synthetic data, providing a practical basis for subsequent demonstrations. The main focus then shifts to converting all character columns to factors using base R and the dplyr package.
We start by exploring the base R method, including identifying character columns, using lapply(), and addressing factor levels. Following the base R section, we look at the dplyr approach. Here, we showcase the use of is.character() and as.factor() within mutate().
Before we conclude the post, we look at the advantages and disadvantages of both methods, aiding in selecting an approach tailored to your workflow. Finally, the conclusion recaps key takeaways, emphasizing the importance of choosing a method aligned with your preferences and project requirements.
Prerequisites
Before getting into the details of converting character columns to factors in R, ensure you have the necessary prerequisites. First and foremost, have R installed on your system. Check your R version using the R.Version() function and, if necessary, update R to the latest version for optimal compatibility.
Additionally, familiarity with basic R functions is beneficial, but do not worry if you are still a novice. This post covers fundamental concepts and functions related to data type conversion, making it accessible for users at various skill levels.
Data Types
Data types play an important role in organizing and interpreting data in R. Common data types include numeric, character, and factor. Numeric data represents quantitative values, while character data encompasses strings of text. Factors, a unique R data type, categorize variables with distinct levels, making them particularly valuable for handling categorical information.
Choosing the appropriate data type is crucial for efficient analysis. The as.character() function converts data to character type, facilitating the manipulation of text-based information. Conversely, as.numeric() transforms data into a numeric format, which is essential for mathematical operations. However, understanding the nature of your data is paramount. We can use the is.factor() function to identify whether a variable is already a factor, providing insights into its structure. Here is an example:
In this post, we will work with the as.factor() function, which proves invaluable for converting character columns into factors. By using these functions, we can ensure that our data is well-suited for the analyses.
Synthetic Data
Here is a dataset to use to practice converting character columns to factors in R:
# Set seed for reproducibility
set.seed(123)
# Generate a cognitive science dataset
n <- 100 # Number of observations
# Variables related to reaction time
reaction_time <- rnorm(n, mean = 500, sd = 50)
stimulus_intensity <- sample(c("Low", "Medium", "High"), n, replace = TRUE)
# Variables related to memory performance
memory_performance <- rnorm(n, mean = 75, sd = 10)
study_material <- sample(c("Text", "Images"), n, replace = TRUE)
# Variables related to participant demographics
participant_age <- rnorm(n, mean = 25, sd = 5)
participant_gender <- sample(c("Male", "Female"), n, replace = TRUE)
# Combine into a data frame
cognitive_data <- data.frame(
Reaction_Time = reaction_time,
Stimulus_Intensity = as.factor(stimulus_intensity),
Memory_Performance = memory_performance,
Study_Material = as.factor(study_material),
Participant_Age = participant_age,
Participant_Gender = as.factor(participant_gender)
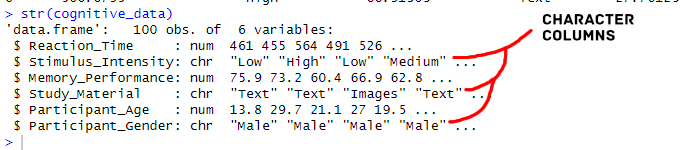
)Code language: R (r)In the code chunk above, we first used set.seed(342) for reproducibility. Next, we created variables such as n, reaction_time, and stimulus_intensity. We used rnorm() for generating reaction time data and sample() to simulate stimulus intensity categories. We then generated memory performance data using rnorm() and determined study material conditions with sample(). Lastly, we created demographic variables like participant_age and participant_gender. The resulting dataframe, cognitive_data, will act as the simulated data from a cognitive science experiment, incorporating factors like reaction time, memory performance, and participant characteristics.
Convert All Character Columns to Factor with Base R
In this section, we will explore the process of converting all character columns to factors using base R functions. This method is effective for users who prefer working with fundamental R functions without relying on external packages.
Identifying All Character Columns
Before converting character columns to factors, it’s crucial to identify which columns are of character type. Next, we can combine the sapply() function with is.character() to identify character columns in a dataset. Combining these functions will give us a logical vector indicating the character columns’ locations.
Using lapply() and as.factor() to Convert all Character Columns to Factor in R
Once character columns are identified, we can use the lapply() function to convert all character columns to factor. Using lapply() allows us to iterate over the identified character columns and apply the as.factor() function to each. Here is an example:
# Identify character columns
char_columns <- sapply(cognitive_data, is.character)
# Use lapply to convert identified character columns to factors
cognitive_data[char_columns] <- lapply(cognitive_data[char_columns],
as.factor)Code language: R (r)In the code chunk above, we integrate the concepts from the previous sections. Initially, we create a logical vector, char_columns, using sapply() to check which columns in the cognitive_data dataset are of character type. This vector serves as a guide for identifying character columns.
The second part of the code uses lapply() to convert the identified character columns to factors systematically.
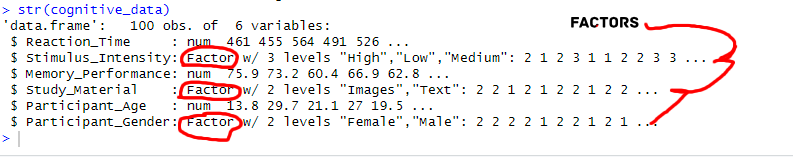
We iterate through each character column by applying the as.factor() function within lapply(), transforming its data type to factor. Using [char_columns] ensures that this conversion is applied explicitly to character columns while leaving others untouched. Note that using [] is a method to select columns in R.
Dealing with Factor Levels
When converting character columns to factors, managing levels is important. Levels represent unique values within a factor column. In scenarios where the dataset has changed or levels need adjustment, the levels() function comes into play. By using levels(), users can view the current levels and modify them as needed, ensuring consistency and accuracy in the factor representation of character columns. Here is an example code of how we can have a look at the levels:
levels(cognitive_data$Stimulus_Intensity)Code language: R (r)We can also use the levels() function to change the factor levels in R.
In summary, the base R method involves identifying character columns, using lapply() for a systematic conversion. This approach provides a fundamental yet powerful technique for handling data type conversions in R.
Convert All Character Columns to Factor in R with dplyr
In this section, we will convert all character columns using dplyr, a powerful package for data manipulation in R. We’ll explore key functions like mutate(), mutate_if(), and the versatile across() along with the helpful is.character() and as.factor() functions. Piping (%>%) will also be employed to streamline our workflow.
Using is.character and as.factor with mutate()
Here is how we can use where(), is.character(), and as.factor() to change all character columns to factor in R:
cognitive_data <- cognitive_data %>%
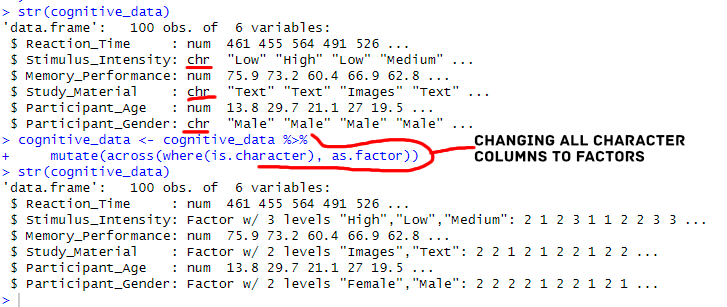
mutate(across(where(is.character), as.factor))Code language: R (r)In this code chunk, we used mutate() in conjunction with across() and is.character() to convert all character columns to factors. Using across() ensures the operation is applied across multiple columns, while is.character() identifies the character columns for transformation. The resulting dataset, cognitive_data, now maintains the desired data type adjustments. Another option is to use the mutate_if() function:
cognitive_data <- cognitive_data %>%
mutate_if(is.character, as.factor)Code language: R (r)Here, mutate_if() simplifies the process by directly targeting character columns based on the specified condition (is.character). The subsequent application of as.factor() ensures a consistent and effective conversion of the identified character columns.
Comparing Methods for Changing Multiple Character Columns to Factors in R
In the base R method, functions like lapply and as.factor() offer simplicity and ease of implementation. However, it may pose challenges when dealing with levels, requiring additional attention to ensure consistent factor levels across converted columns. On the other hand, the dplyr mutate method provides a concise and expressive syntax, especially when combined with functions like across() and mutate_if. Its drawback lies in the dependency on the Tidyverse, which might only be suitable for some workflows.
The choice between base R and dplyr methods depends on your familiarity with these tools and the specific requirements of the data analysis. The base R method might be preferable if you are comfortable with base R functions and seeking a straightforward approach. Conversely, individuals working within the Tidyverse ecosystem may find the dplyr mutate method more coherent and integrated into their data manipulation workflows. Factors like code readability, maintainability, and project dependencies will guide selecting the most suitable approach.
Conclusion
In conclusion, this tutorial explored two distinct methods for converting all character columns to factors in R, offering flexibility based on your preferred workflow. The base R approach, employing functions like lapply and as.factor(), caters to simplicity, making it accessible for users accustomed to base R functions. On the other hand, the dplyr mutate method provides a concise and expressive alternative, particularly for those working within the Tidyverse ecosystem.
Key takeaways include the importance of understanding different data types in R, choosing appropriate methods based on your workflow.
As you navigate your data analysis journey, choose the method that seamlessly aligns with your preferences and project requirements. Whether you choose the familiarity of base R or the expressive syntax of dplyr, this guide equips you with the tools to manage character columns efficiently.
If you found this tutorial valuable, consider sharing it on social media. Feel free to comment below with any suggestions, requests, or insights. If you use these methods in your reports or papers, referencing this post will be greatly appreciated.
Resources
- Not in R: Elevating Data Filtering & Selection Skills with dplyr
- How to Sum Rows in R: Master Summing Specific Rows with dplyr
- Countif function in R with Base and dplyr
- Sum Across Columns in R – dplyr & base
- How to Standardize Data in R with scale() & dplyr
