Table of Contents
PsychoPy
This post will describe why I prefer using PsychoPy before other software. When I started getting involved in research (i.e., when doing my Bachelor’s and Masters’s theses), I used the software that was accessible to me. That is, some of the most commonly used software in the department I was studying at. After starting my Ph.D. studies, I got increasingly annoyed with one of the software we used to build experimental tasks in (i.e., E-prime).
First, I was unfamiliar with the scripting language (called ‘e-basic’) that was needed, for instance, to solve pseudo-randomization and send signals to a specially built device I was going to use in my experiments. Second, I prefer to use Linux systems, and e-prime is limited to Windows. In fact, e-prime seemed to work best with Windows Vista or earlier, but our lab computers were, and are, running Windows 7 (which might have changed back then). Some time into my Ph.D. studies, I found PsychoPy, which is written in Python, a programming language that I did know. It seemed like a good alternative to e-prime, and other similar software, to create experiments in.
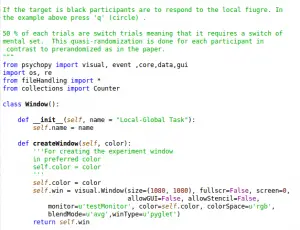
PsychoPy’s website states that it is an open-source application that presents stimuli and collects data in experiments. Being Open Source also means that the software is free. Importantly, it can be run on Linux, OS-X, and Windows. One of the features that I like most is that I can choose to write code or to use an experimental builder such as e-prime. I prefer to write scripts and not use a Builder. However, when coding existing modules and methods in PsychoPy.
Since I am running Linux on my computers, I can use my favorite Python IDE (Integrated Development Environment), Spyder. I import the modules and/or methods I need in my script, and then I can run experiments from within Spyder. I assume, but don’t know, that you can use your favorite IDE in other OS’ also… PyCharm, however, is also one of the best Python IDEs I have used.
Why students should use PsychoPy
For the less coding-experienced people, there is also the builder view. An experimental builder is a graphical interface with menus and, often, drag-and-drop-like features.
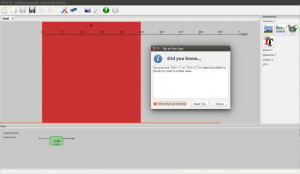
The builder view is one of the reasons I have started to suggest students that I meet use PsychoPy to build experiments for their projects and thesis. Another reason is that Python as a scripting language is also great because the cognitive science students at our department take at least 15 ECTS in Python programming. It might be worth mentioning that since Python is a popular programming language with a great and helpful online community (see Stackoverflow tags “#Python” and “#Psychopy“) . Finally, because the students can install it on their own computers, they can sit at home, in the library, or wherever they want to complete their experiments.
Installing PsychoPy
If you are interested in installing it from Ubuntu, you can add the Neuro Debian repository:
wget -O- http://neuro.debian.net/lists/trusty.de-m.full
sudo tee /etc/apt/sources.list.d/neurodebian.sources.list
sudo apt-key adv --recv-keys --keyserver hkp://pgp.mit.edu:80 0xA5D32F012649A5A9
sudo apt-get update && sudo apt-get install psychopyCode language: Bash (bash)Another way to install it is to use Pip;
pip install psychopyCode language: Bash (bash)Note that you might need to install some dependencies (i.e., some other Python packages) to get it to run under Ubuntu (it may prove useful to check the post linked below).
If you are running Linux Mint Debian Edition (LMDE) you can try to use the instructions in an earlier post How to Install PsychoPy on LMDE.
PsychoPy Resources
As far as I know, there is only one book that mentions and gives an example of how to use PsychoPy: Computational Neuroscience and Cognitive Modelling: A Student’s Introduction to Methods and Procedures. If anyone reads this and knows about other books, please leave a comment with the title and author’s name.
You can also look at my example script: SART written in PsychoPy. Finally, here you will find some useful links to Python (and R) related resources: Python Resources. If you want to learn how to make a Psychomotor vigilance test in PsychoPy or download it, check that post out.
References
Peirce, J. W. (2007). PsychoPy-Psychophysics software in Python. Journal of Neuroscience Methods, 162(1-2), 8–13. Retrieved from https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2018741/

Actually, there are more books now, and with more depth than the one you link to
– “Building Experiments in PsychoPy” by Peirce and MacAskill is the most in-depth covering the Builder interface.
– “Programming Visual Illusions for Everyone” by Marco Bertamimi is great, fun way to learn how to use PsychoPy stimuli from Python code.
– “Python for Experimental Psychologists” shows you how to create experiments in either PsychoPy or PyGaze, although it’s written using Python2 syntax which I wouldn’t recommend now (not fully compatible with Python3).
Hey Jonathan,
Yes! I actually own “Building Experiments in PsychoPy” but I have to find time to read it. I’ll update this post, with these books, during the weekend.